Pair Correlation Analysis¶
Contents
The pair correlation analysis window can be opened from the PAM main window by clicking on Advanced Analysis -> PCF Analysis or from the command window by typing in PCFAnalysis.
The pair correlation method ([Digman2009]) extracts information about the spatial behavior of molecular movement using rapid line-scanning and a correlation function between pairs of spatial locations in the sample. It is especially useful to measure (spatially and directionally resolved) diffusion and transport of molecules across barriers.
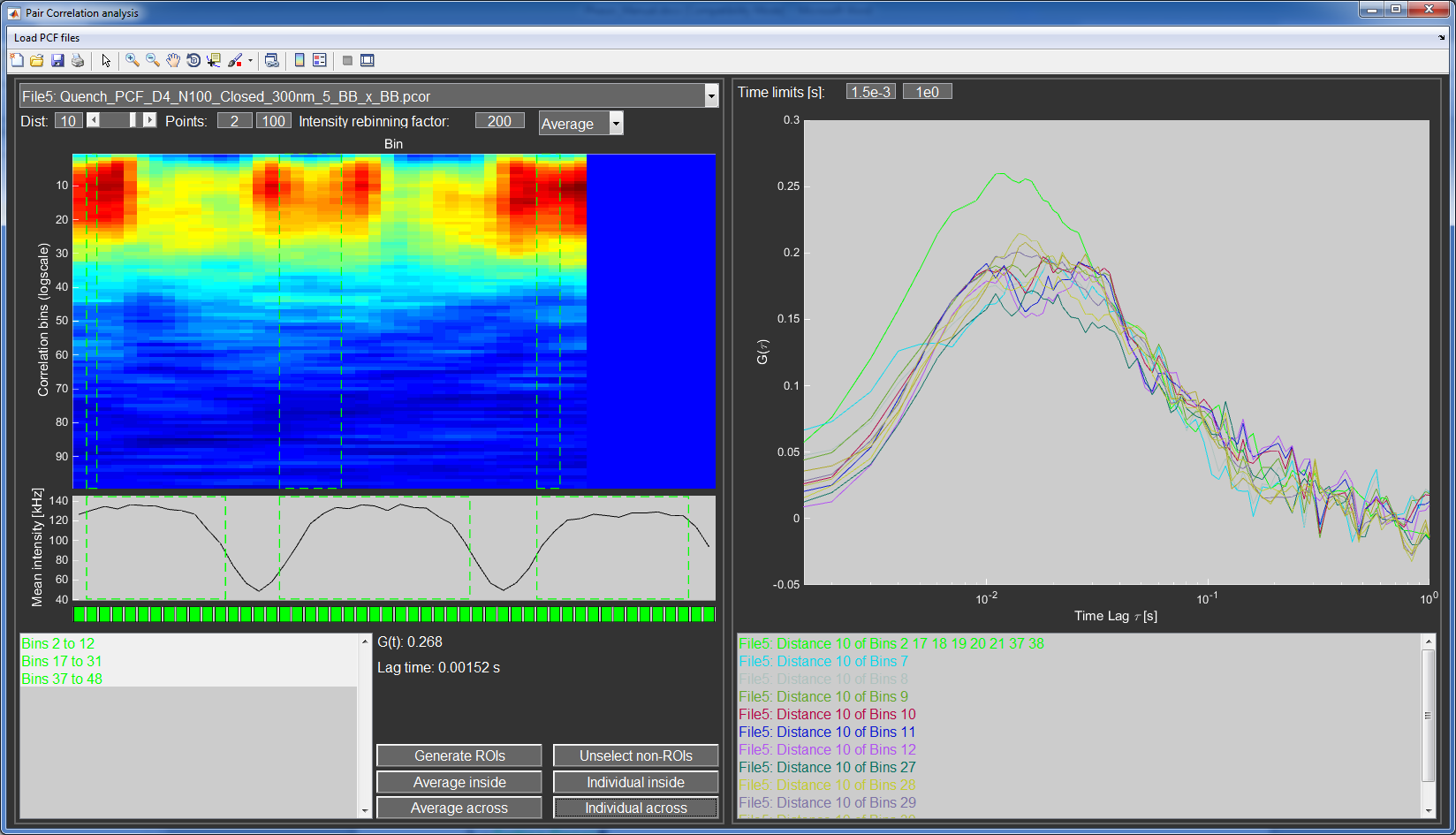
The PCF Analysis window is divided into two main parts. On the left side the PCF carpet is displayed and the user can select regions and extract individual correlation curves. The right side is reserved for display of the individual correlation curves that can be averaged and exported.
Data handling and loading data¶
Currently, the program only supports files created with PAM with the file ending .pcor. These files are based on standard .mat files and can also be accessed in the MATLAB command window.
To load new files, got to Load PCF files and select either Load New File or Add Files. Load New File removes previously loaded files, while Add Files adds the new one to the old ones. The currently selected file can be selected with a popupmenu. The loaded data are stored in the global variable PCFData.
PCF Carpet display¶
Pair correlation data is a 4 dimensional matrix (position, time lag, distance lag, and backwards and forwards correlation). This makes it very hard to properly display the data in a meaningful way. In this program a carpet for a single distance lag is displayed, with the laser position (=spatial bin) as the horizontal axis and the time lags (as a quasi-logarithmic scale) as the vertical axis. The correlation amplitude is colorcoded using the Jet colormap.
The user can select the displayed spatial lag with the Dist editbox and slider. Furthermore, the editboxes for Points restrict the temporal lags displayed, useful to remove artifacts at early or late time points. Initially, the average of the forward and backwards correlation is plotted (to increase S/N). In cases where diffusion might be direction dependent, the user can select to only display and analyze the forward or backward correlations with a popupmenu.
Right clicking the carpet gives the user additional options:
- Normalize: Normalizes the carpet to the maximal value for each spatial bin. Selecting it again replots the original data.
- Plot: This option allows the user to switch between the correlation, an intensity or an arrival time carpet.
- Export: This exports the currently plotted carpet to a new MATLAB figure.
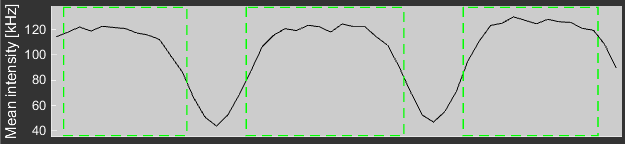
Below the carpet is a plot displaying the mean intensity profile of the scanned region. With a right click the user can switch to the mean arrival time profile.
Selection of bins and ROIs¶
In most cases not all bins of the carpet are of particular interest for further analysis (e.g. parts outside of a scanned cell) or the bins need to be sorted into different categories.
Individual bins can be unselected with the green boxes directly below the intensity profile plot. A right click unselects a single bin (indicated by a red box). Note that each spatial bin is part of two correlation curves and both are omitted. A second click selects it again. The left mouse button toggles the selection for all bins at once. All further analysis and averaging require the definition of regions of interest (ROIs). ROIs can be generated in several different ways: * Generate ROIs Button: Clicking this button will select a ROI for each consecutive stretch of active bins. That means that if all bins are active a single ROI spanning the full range will be selected. * Left Click in the Intensity Profile: Left clicking and dragging the mouse in the intensity profile plot selects the area as a single ROI (even across inactive bins) * Left Click in the Carpet: Left clicking and dragging the mouse in the carpet also selects a ROI. However, due to the spatial lag, this will generate a larger ROI, since it has to include both spatial bins used for the correlation.
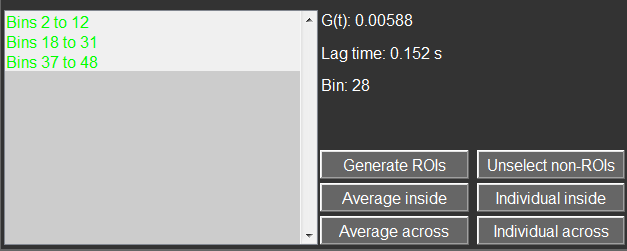
The individual ROIs are shown in a listbox at the bottom left. They can be repositioned using the arrow keys and resized with the ´+´ and ´-´ keys. The standard color of a ROI is green. Selecting them and pressing the c key allows the user to select a new color.
Curve extraction and averaging¶
Once the ROIs are defined, the user has to extract the relevant bins either as individual or averaged correlation curves to the correlation curve part on the right side of the window. Hereby, the program distinguishes between two different types of curves:
- Both spatial bins of a particular correlation curve are situated within a single ROI. This case is called “inside�? and these curves can be extracted with the Individual inside button as an entry for each single one or as a single averaged curve with the Averaged inside button or the enter/return key. The individual curves get a random color while the averaged curve has the average color of the ROIs.
- When the first and second bins of a correlation curve are situated in two different ROIs, the case is called “across�?. This requires that at least two ROIs are selected. These curves can be extracted with the Individual across button as an entry for each single one or as a single averaged curve with the Averaged across button or the enter/return key. The individual curves get a random color while the averaged curve has the inverse of the average color of the ROIs (1-RGB).
Individual curve display and exporting¶
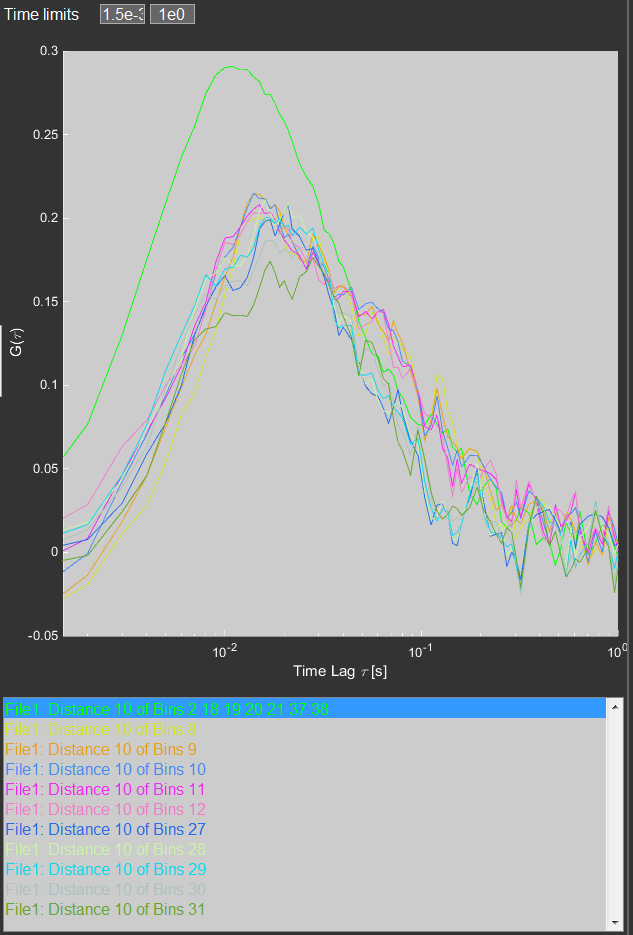
The individual and averaged curves extracted from the ROIs are displayed and listed on the right side of the program.
The arrow keys can toggle the display of the selected curves. A further averaging of the curves can be achieved by selecting them and pressing the ´+´ key.
Pressing enter/return will export each selected curve as a .mcor file that can be loaded and fit with the FCSFit program.
| [Digman2009] | Digman, M.A. Imaging barriers to diffusion by pair correlation functions. Biophys. J. 97(2), 665-673, (2009). |